A Silent Crisis
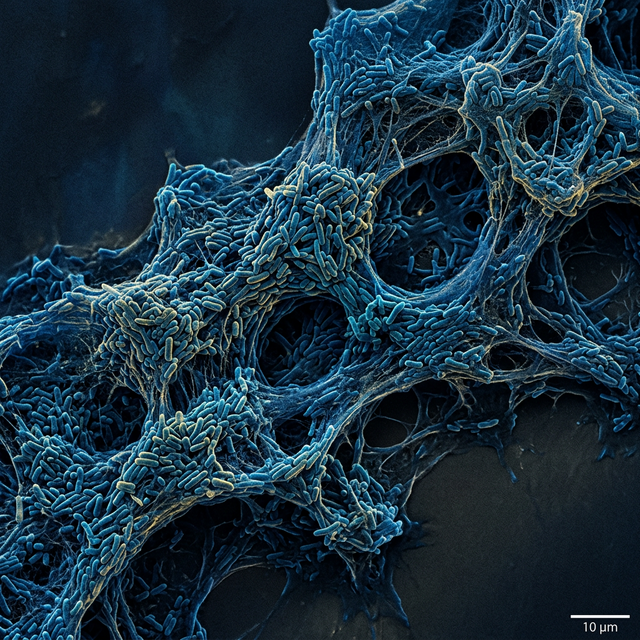
Current antibiotics often fail not because bacteria are genetically resistant, but because they tolerate stresses. In chronic conditions like Cystic Fibrosis and COPD, pathogens like M. abscessus and P. aeruginosa modify cell physiological properties to optimize tolerance to host immunity and conventional antibiotics.
The approach taken to develop new antibiotics has focused almost exclusively on targeting discrete molecular mechanisms that are believed to be essential systems for life and virulence. However, evolution and adaptation readily permit bacteria to circumvent such chemotherapeutic strategies. We therefore propose a new paradigm for drug discovery, one that bacteria cannot overcome to cheat death.
- Targeting global markets in health, nutrition, and industry
- Discovery of compounds driving mechanical change
- Harnessing mechanical change in evolutionarily disparate species